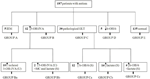
Typical incompatibilities and inconsistencies in genome-scale models and databases. A key bottleneck in the pace of reconstruction of new high quality metabolic models is our inability to directly make use of metabolite/reaction information from biological databases (e.g., BRENDA, KEGG, MetaCyc, EcoCyc, BioCyc, BKM-react, UM-BBD, , Rhea, PubChem, ChEBI etc.) or other models due to incompatibilities of representation, duplications and errors, as illustrated in Figure Figure1 1. Already over 75 genome-scale models are in place for eukaryotic, prokaryotic and archaeal species and are becoming indispensable for computationally driving engineering interventions in microbial strains for targeted overproductions, elucidating the organizing principles of metabolism and even pinpointing drug targets. These reconstructions account for reaction stoichiometry and directionality, gene to protein to reaction associations, organelle reaction localization, transporter information, transcriptional regulation and biomass composition. Increasingly, metabolite and reaction information is organized in the form of community, organism, or even tissue-specific genome-scale metabolic reconstructions.

The ever accelerating pace of DNA sequencing and annotation information generation is spearheading the global inventorying of metabolic functions across all kingdoms of life.
